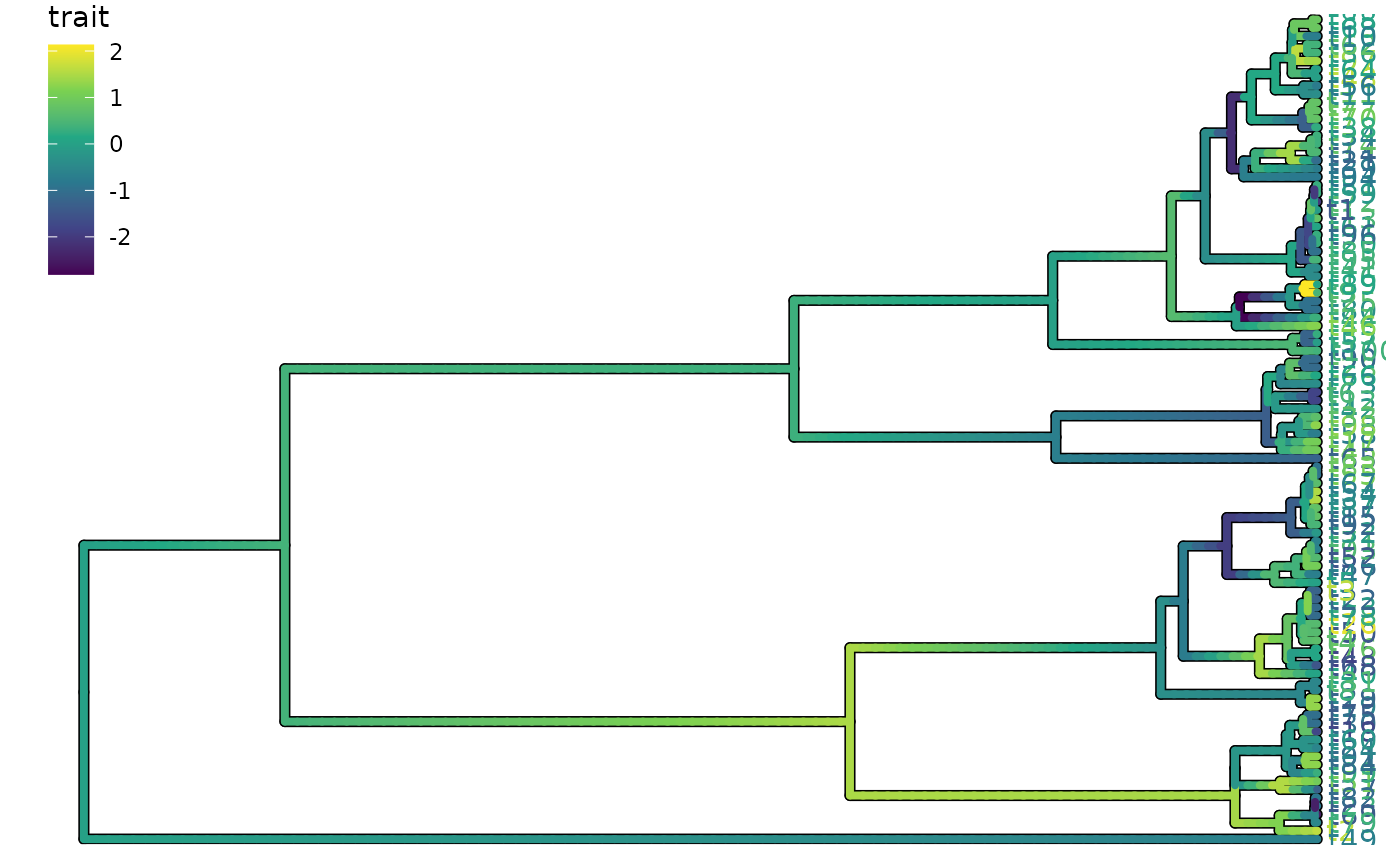
Make an automatic ggplot2 plot for a pf object
autoplot.pf.RdMake an automatic ggplot2 plot for a pf object
Usage
# S3 method for class 'pf'
autoplot(
object,
columns = NULL,
layout = "circular",
suppress_tiplabels = FALSE,
suppress_tippoints = FALSE,
edge_traits = FALSE,
continuous = "colour",
size = 1.4,
outline_size = 1.4 * size,
...
)Arguments
- object
A
pfobject to plot- columns
Columns to plot along with the phylogeny. Can use bare column names or any other
tidyselectsyntax- layout
ggtree::ggtree()layout to use.- suppress_tiplabels
If
TRUE, don't draw tip labels.- edge_traits
A logical indicating whether the continuous avriable refers to edge traits, where the edge is determines by the terminal node. (default is FALSE, which means the variable refers to a node trait).
- continuous
continuous transition for selected aesthethic ('size' or 'color'('colour')). It should be one of 'color' (or 'colour'), 'size', 'all' and 'none', default is 'colour'.
- ...
Other arguments passed to or from other methods.